I'm searching for a solution to automatically generate 2D molecule representation from different sources (SDF for example).
No graphical interface, library would be a plus.
I'm searching for a solution to automatically generate 2D molecule representation from different sources (SDF for example).
No graphical interface, library would be a plus.
Arguably the best depiction software available is the 2D layouter integrated in the Cactvs toolkit (see www.xemistry.com/academic for free academic/educational versions). It is used for example in PubChem and by various 3rd-party software vendors (IDBS, Optibrium, etc.).
It provides both coordinate generation and depiction (via multiple renderers, including pixel images [GIF,PNG,bitmaps], SVG, WMF, EMF, PICT, EPS, PDF, MIF, SWF). Full stereochemistry support, including stereo-centers with free electrons, odd and even allenes, square planar stereochemistry - and proper trans double bonds in macrocycles - as far as I know, it is the only layouter which can do that. It knows about standard ring system orientations - so a steroid will always have its well-known appearance, regardless of the order of the input atoms, and pyridine etc. come out with the hetero atom at the bottom, not somewhere random, etc.. Aligned layout on a common substructure fragment is supported, and DNA/RNA/Peptide polymers are automatically recognized and laid out in a proper chain sequence. There is also a basic reaction layouter.
The toolkit has an easily programmed script interpreter, so setting up a batch script with custom parameters is simple. Over 50 structure file formats are automatically recognized and supported, including SDF.
All major platforms (Windows, OSX, Linux, Solaris, FreeBSD) are supported. We sell turnkey solutions for these problems, too, but these are limited to commercial customers. Link libraries for integration into custom software may be licensed.
Neutral comparisons:
The Chemistry Development Kit can do this with the StructureDiagramGenerator. A full code example can be found in this blog post. The basic use looks like:</p>
StructureDiagramGenerator sdg = new StructureDiagramGenerator();
sdg.setMolecule(someMolecule);
sdg.generateCoordinates();
Molecule layedOutMol = sdg.getMolecule();
If your input already has 2D coordinates, then the problem is much simpler, and you only need depiction. Then, you can use the rendering functionality from the CDK. Several tutorial posts can be found as links in this README. This will graphics like:
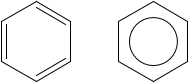
I wrote a XSLT stylesheet to transform PC-Compound/XML to SVG. See my blog here
xsltproc --stringparam scale 30 --stringparam xradius 2 \
--stringparam show-bounds false \
src/xsl/pubchem2svg.xsl \
CID_16204538.xml > file.svg
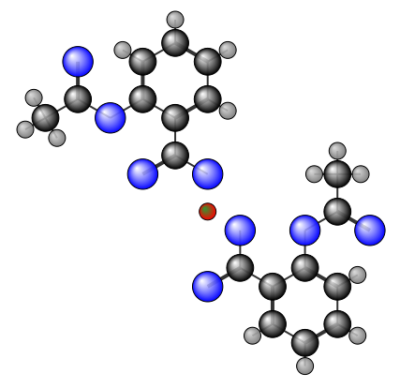
There's a simple tool called mol2ps / mol2png which you can call from the command line.
OpenEye's OEDepict library would be a perfect solution for this problem. OEDepict offers rapid 2D coordinate generation and artful rendering of chemical structures. 2D coordinates can be generated from SMILES or from 3D structures. Multiple image file formats are supported including but not limited to: PNG, SVG, PS, EPS, and PDF. In addition to handling molecule 2D coordinate generation and depiction routines, OEDepict provides a powerful engine that supports:
The library is written in C++ but has bindings for Python, Java, and .NET. Most standard platforms are supported including Microsoft Windows, Mac OS X, and a wide variety of Linux flavors.
OEDepict is free for non-commercial use and is used widely throughout the pharmaceutical industry and embedded in many third-party applications.
OEDepict also has a companion library, Grapheme TK that provides additional mechanisms to enhance standard 2D depictions based on 3D properties. I've included a few example images below.


Use of this site constitutes acceptance of our User Agreement and Privacy Policy.
2D coordinate generation or depiction or both?
Yeah, I originally assumed generation of 2D coordinates was meant, but lost confidence in that... clarification most welcome.
http://cdk.sourceforge.net/