Entering edit mode
4.6 years ago
deepak18
▴
10
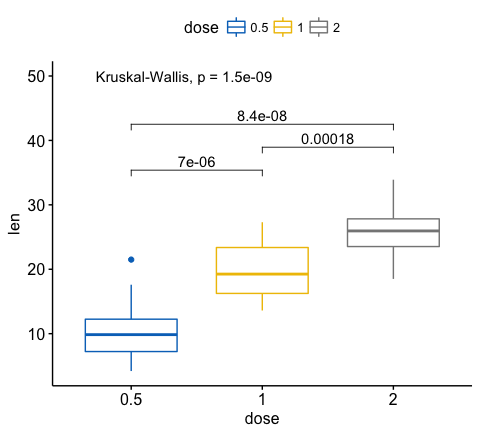
I have Isoform level RNA-seq data from The Cancer Genome Atlas and I want to check the difference the difference in the level of a particular isoform between cancer and normal samples. Is there any software or R package which I can use to plot a boxplot showing difference in that isoform between normal and cancer sample with p-value?


If I may ask, how did you obtain Isoform level RNA-seq data from TCGA? I am trying to do so via TCGAbiolinks R package, however, without positive results