Entering edit mode
4.2 years ago
zhoujie
•
0
Recently I received some single cell RNA-seq data from a producer. The data format is like this:
I1: sample barcode R1: 150bp reads R2: 150bp reads
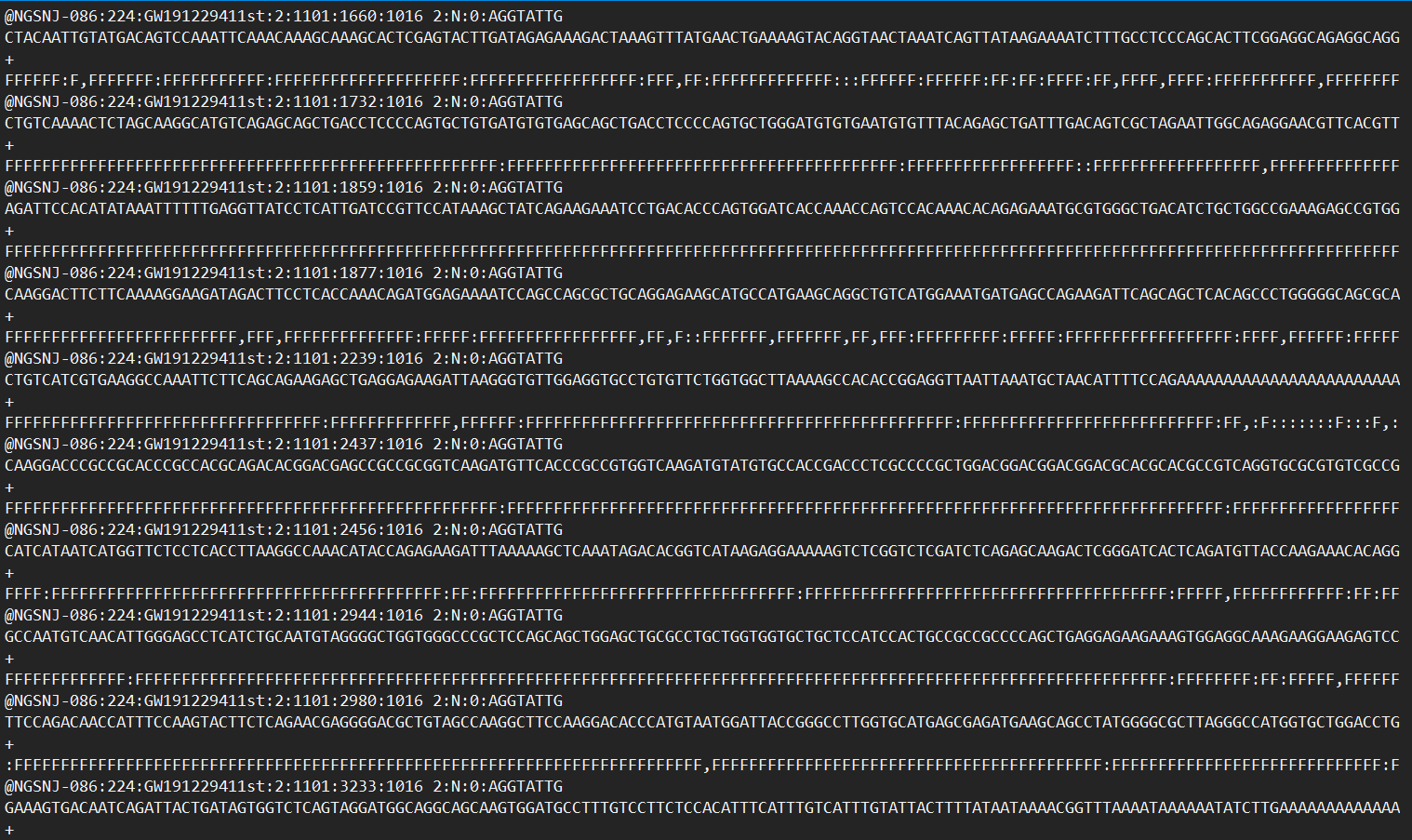

Can someone teach me how to analyze this kind of single-cell RNAseq data? I don’t know how to put it into the Cellranger.


Did you read the cellranger manual? The 10x website? Did you ask the software what it expects in the command line? Did you try the test data that comes with cellranger?
Yes I read the cellranger manual and the expected format should be like 28*91bp for R1 and R2. I have run it by cellranger, but what I got is only 400 cells, which is much less than expected. So I wonder if it is some preprocess thing I need to do first.
You just give cellranger the fastqs as they are. Just like the website says.
So my question is how to transform the format to the expected format?
What format? Cellranger will not be confused by longer reads.
Yes but the result is bad. There is no cell barcode in the fastq file.
Define 'bad', please? What do you want to do?
https://github.com/2kjiejie/30-297112163/blob/master/631581549586_.pic.jpg https://github.com/2kjiejie/30-297112163/blob/master/621581549580_.pic.jpg
Here are the screenshots for R1 and R2