Many experts claim that the negative binomial distribution is better than the Poisson distribution for modeling discrete RNA-Seq data.
Is this because
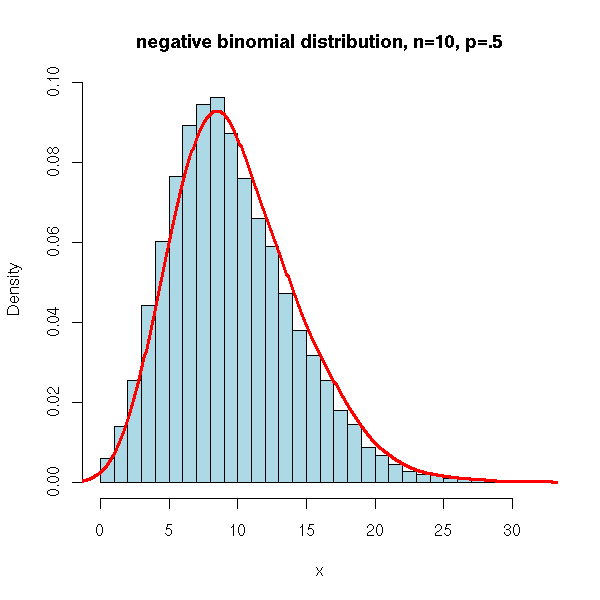
- The expression profiles of organisms resemble the negative binomial distribution - i.e.if I binned real gene transcription counts from an RNA-Seq experiment it would look like the plot below.
or
- The sampling error of gene expression is such that the true population mean of a gene looks like the negative binomial - i.e the true mean expression level of that gene is probably more (because of the skew) than the mean expression level of a sample of reads of that gene drawn from replicates.
or are these two concepts the same thing?


Moved to separate question per suggestion
Please don't append questions to other questions, I think your questions would stand quite well on their own as a separate topic. Please ask a separate question.