Hi,
I am new to RNA sequencing and I am a bit confused with the HT-Seq read count options and I want to know whether I am thinking in the right direction. I have a set of paired-end strand-specific RNA-Seq reads and I am now trying to count the reads in a set of features (genes).
The HT-Seq documentation says that the option "stranded" by default is set to "yes" which means that HT-Seq assumes the reads to be strand-specific. They also say
"If your RNA-Seq data has not been made with a strand-specific protocol, this causes half of the reads to be lost. Hence, make sure to set the option --stranded=no unless you have strand-specific data! "
This makes sense, since if I use "stranded=yes" option for non-strand specific data, the reads mapping to the opposite strand of the feature will NOT be counted.
However, this makes me wonder, if I use "stranded=no" even for strand-specific data, it would not affect my counts in any way. Is that correct ? Because with "stranded=no", it does not matter if a read maps to the same or the opposite strand as the feature. It would be counted as long as it is mapping to a feature, regardless of the strand.
So then a follow up question comes to mind as to why HT-Seq even has the "stranded=yes" and "stranded=reverse" options.
I am sorry if this is a very naive and incorrect question, but I really need to get the strand-specific concept clear in my mind.
Any help would be much appreciated.

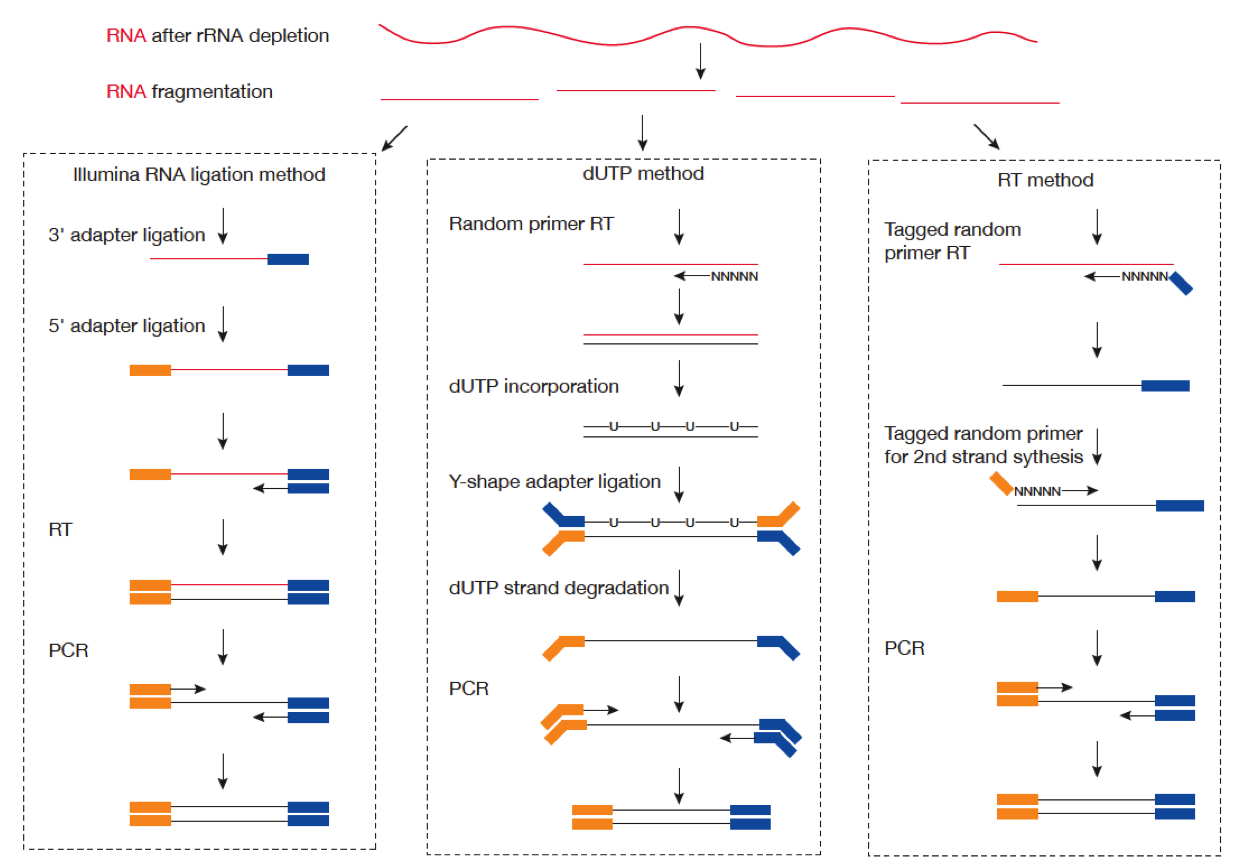 Image Credit: Zhao Zhang
Image Credit: Zhao Zhang