I developed a cox model based on rna-seq datasets. To assess the performance of the model, I applied model to patients and calculated a risk score for every patients. Then we divided the patients into a high risk and low risk group (the median value of all risk score was cutoff value). I applied KM analysis (log-rank test) to these 2 group. Next, I applied the ROC analysis to the model.
In my opinion, I believed a good model should have a log-rank test P-value < 0.05 and a large AUC value of ROC analysis (such as >0.70).
However, the results were pretty wired and I did not know how to explain them. The P-value of log-rank test were more than 0.05, but the AUC value of ROC was more than 0.70. The figures could be seen as follow.
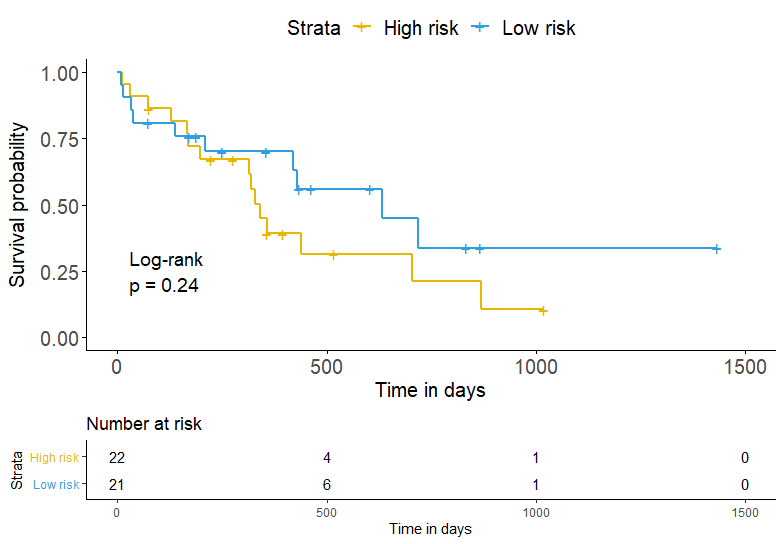
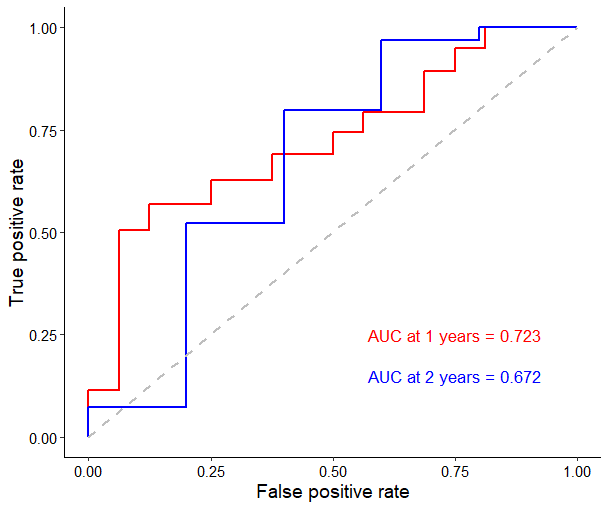
Is there somebody who can help me understand this result?? I really need your help. Many thanks!!


I am not sure that I understand what you're doing but assuming that the ROC analysis is about predicting survival at 1, resp. 2 years, then it is not doing the same thing as the logrank test. With the ROC analysis you're testing prediction of survival at a particular time point for each group separately, with the logrank test you're comparing the survival distributions between two groups. If the survival distributions differ significantly at 5 years but not at one year then your model shouldn't predict survival at one year very well but the logrank test should still pick up a difference since it considers the distributions (i.e. at all time points). I also think that your assumption of good p-value being linked to good AUC relies on the mistake that p-values are indicative of the strength of an effect. There are plenty of cases where a detectable effect doesn't translate into a strong statistical signal. Although what constitutes a good AUC value depends on the context, I would generally consider AUC below 0.8 to be not very good especially in a medical context where false positives can be costly. Finally, it seems likely that your analysis is underpowered and you may need more patients.
Thanks for your guidance. It gives me a lot of inspiration. The size of my patients was indeed not large enough. I will collect more patients in the future. However, I still have a question: what should I do to assess the performance of a model comprehensively? Is it ok to apply ROC analysis and logrank test? Or there are other methods I can try? I would be very grateful if you can give me some suggestions.