RNA-seq Bioinformatics: A Practical Introduction
Mapping, Visualization, Basic Analyses

When? October 16th - 17th 2014
Where? Leipzig, Germany
Scope and Topics
The purpose of this workshop is to get a deeper understanding in High-Throughput Sequencing (HTS) with a special focus on bioinformatics issues. Advantages and disadvantages of current sequencing technologies and their implications on data analysis will be discovered. The participants will be trained on understanding their own HTS data, finding potential problems/errors therein and finally begin writing their own analysis pipelines. In the course we will use a real-life RNA-seq dataset from the current market leader illumina.
By the end of this workshop the participants will:
- be familiar with the sequencing methods from various platforms:
- Illumina
- 454 (Roche)
- Ion Torrent
- PacBio
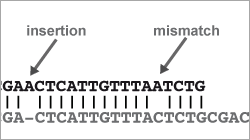
- be aware of the different error sources of these machines
- understand common data formats and standards
- know relevant tools for HTS data processing
- automate tasks with shell scripting to create reusable data pipelines
- perform basic analyses (gene quantification, differential expression)
- plot and visualize results
- be able to continue and reuse all analyses
Requirements
- basic linux knowledge (shell usage, common commands). You should be familiar with the commands covered in the Learning the Shell Tutorial
- basic understanding of molecular biology (DNA, RNA, gene expression, PCR, ...)
Target Audience
- biologists or data analysts with no or little experience in analyzing HTS data
Included in the Course
- Course materials
- Catering
- Conference Dinner
Program
- Introduction to sequencing technologies from a data analysts view
- Mechanisms of instruments: Illumina, 454, IonTorrent and PacBio
- Sequencing protocols (mRNA-seq, micro RNAseq, ...)
- Capabilities & limitations: error sources, biases & beyond
- Sequence file formats (fastq)
- Preparation of raw reads: quality control, adapter clipping
- Read mapping
- Alignment methods
- Mapping tools, mapping to a reference genome
- Mapping output
- File formats (SAM/BAM)
- samtools
- bedtools
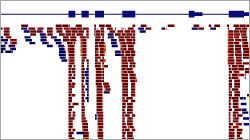
- Visualization of mapped reads
- UCSC: data format for upload
- IGV (local)
- Gene expression quantification (coverage, FPKM)
- Differential expression analysis (with DESeq)
Speakers
David Langenberger (ecSeq Bioinformatics) started working with small non-coding RNAs in 2006. Since 2009 he uses HTS technolgies to investigate these short regulatory RNAs as well as other targets. He has been part of several large HTS projects, for example the International Cancer Genome Consortium (ICGC).
Mario Fasold (ecSeq Bioinformatics) works in the analysis of microarray data since 2007 and developed several bioinformatics tools such as the Bioconductor package AffyRNADegradation and the Larpack program package. Since 2011 he specialized in the field of HTS data analysis and helped analysing sequecing data of several large consortium projects.
Guest-speaker:
Invited speaker (Illumina) will give a short presentation about state-of-the-art sequencing technologies from Illumina.
Key Dates
Opening Date of Registration: April 1st 2014 Closing Date of Registration: August 1st 2014 Workshop: October 16th-17th 2014 (8am - 5pm)
Attendance
Location: Leipzig, Germany. Language: English Available seats: 20 (first-come, first-served)
Registration fees:
- industry rate: 850 EUR
- academic rate: 600 EUR
Travel expenses and accommodation are not covered by the registration fee.
Note: Combine this workshop with our other workshops and get 10% discount.
Contact
ecSeq Bioinformatics
Brandvorwerkstr.43
04275 Leipzig
Germany
Email: events@ecSeq.com
Downloads
---> Register
When you register for this workshop you are agreeing with our Workshop Terms and Conditions. Please read them before you register.
| **sponsored by** | **media partner** | ||
 |
 |
 |





