Hello All,
I would like to ask for advice on using limma for differential expression analysis of scRNAseq data. I usually use Seurat, but I would really like to know whether limma is also a suitable tool for scRNAseq data. I've tried to search for papers and I couldn't find a stable answer to this question.
Thank you very much for your help!
Regards, Anna

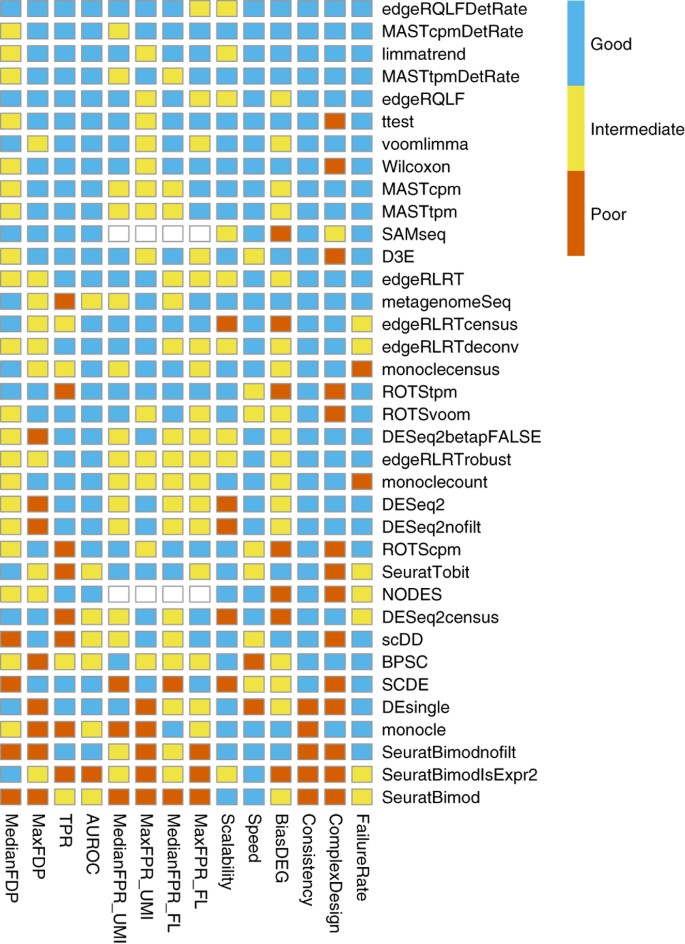

Thank you very much! I will read that paper and will look at the code - I like limma because it is very versatile and allows to perform various comparisons with very variable designs. I will be glad to know that I can use limma for scRNAseq as well.