What is the difference between a Read and a Fragment in RNA-seq, also are the terms single-end and paired-end related?
This diagram illustrates the difference:
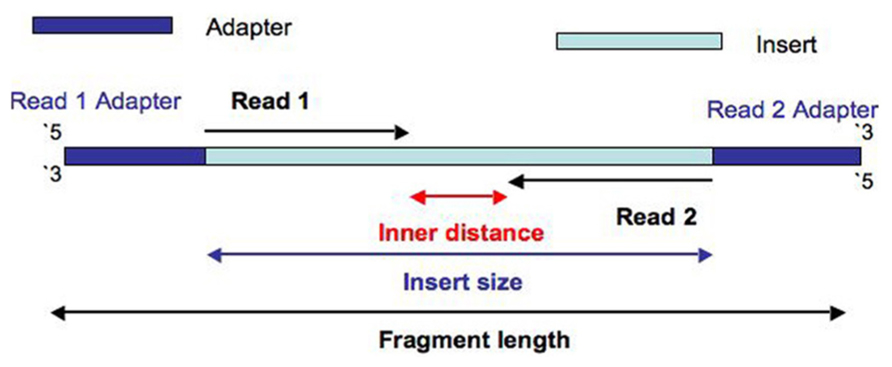
This video of the Illumina sequencing process gives you the context you need to understand what is going on in the diagram.
The difference between fragment and read is the same for RNA-seq, whole genome sequencing, exome sequencing, etc. But there are some terms like 'FPKM' and 'RPKM' where there is additional meaning in an RNA-seq context.
Also, explained in the video above. A paired-end read is one where sequence has been read (from outside inwards off of a sequencing adaptor) and from both ends of a fragment. Since they are generated from the same physical fragment1, they are said to be paired. A single-end read may be generated from only one end of the same fragment.
- 1 or more precisely in the case of Illumina from a cluster of identical2 PCR amplicons of a fragment,
- 2 strictly speaking these amplicons will not all be identical due enzyme introduced errors (hopefully random errors)
Related post: Insert Size And Fragment Size ?
Nice blog/tutorial on the topic: http://thegenomefactory.blogspot.com/2013/08/paired-end-read-confusion-library.html
Use of this site constitutes acceptance of our User Agreement and Privacy Policy.


Have you bothered to read anything describing how Illumina sequencing works?
I read about that, and I found some used Read for single end and fragment for paired end so is it right!!
Not exactly, no.
so what are those mean?