Okay, so I finally completed this task by doing the following:
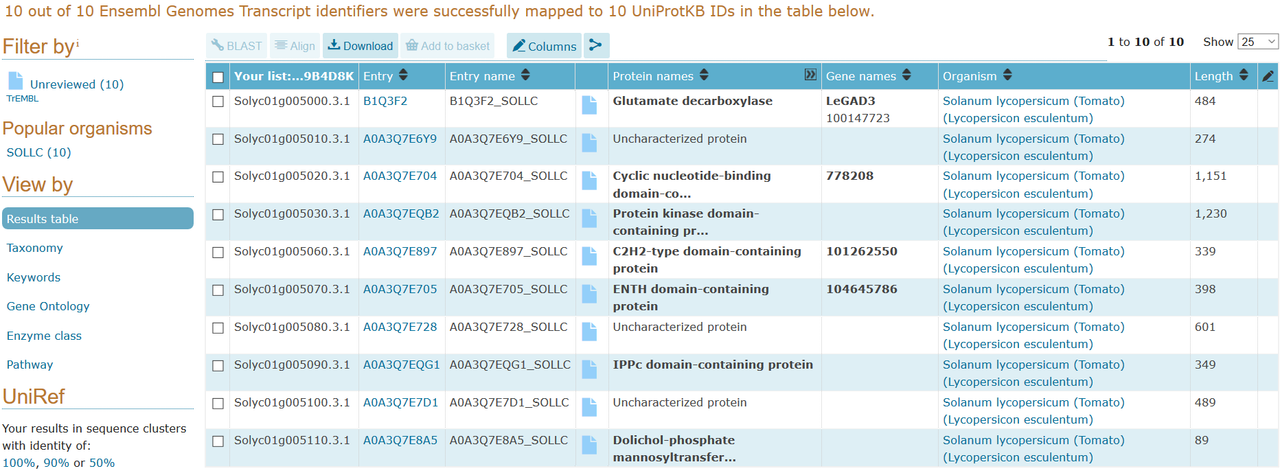
I contacted the Boyce Thompson Institute who ran the Sol Genomics website to see if they had any conversion sheets from their Sol genomics IDs to something more common. They sent me a very good conversion sheet of Sol gene IDs to UniProt IDs.
If someone knows how to share files on biostar I'll attach the conversion sheet to my answer. It's 1.7 MB, so not too big. I can't find the same file in the Sol genomics ftp directory, but I haven't looked that hard either. so if someone knows how to share files like this or if the file is on the sol genomics ftp directory, any help would be appreciated.
So I got my sol gene IDs, got the sol to Uniprot IDs sheet, and used Galaxy to convert the sequences with the info on the conversion sheet.
Most gene enrichment analysis programs take the UniProt IDs, so this conversion did what I wanted it to do.
Also this web based program can do Gene Enrichment analysis using the Sol gene IDs: http://bioinfo.bti.cornell.edu/tool/MetGenMAP
Another good program that can use the Sol gene IDs for gene enrichment analysis: http://mapman.gabipd.org/web/guest/home
So that's how I solved my issue. Thanks for the help guys.


Do you know which Sol Genomics IDs correspond to which GeneID (like in a excell file or something)? Because if that is the case, then i can produce a simple script that does the translation.
If you are actually looking to query several databases than i'm sorry to say i can't help you.
I can't find a chart with GeneID conversions. I'm combing the Sol Genomics website, but can't find it. I know that the tomato genome data is public and on several databases. If anyone has done this before or has the file please share the link. Right now typing in my keywords on google is yielding no results
There is this folder with ID conversion files: ftp://ftp.solgenomics.net/genomes/Solanum_lycopersicum/id_conversion/
Which ID's do you have?
Oh wow, there it is. I have the SolyC IDs that look like Solyc09g009710.2.1.
Unfortunately there does not seem to be any easy way (or a file) to easily translate those to NCBI ID's.
This is exactly what I am looking for. Amazing! Thank you. Is the link updated by the SOL genomics team or are the genes valid for the genome of 2 years ago?
Hi, Could you please send me the tomato conversion sheet on my email: marek_koter (at) sggw.pl ? Thanks a lot, Marek D. Koter
Hello, Kevluv93! I can not convert my Solyc to NCBI-gene ID. I would be grateful if you could send me the excel file to to my email (jamille_agronomia@yahoo.com.br). Thank you very much!
Please, could you also send to me the files with conversion of Solyc numbers to uniprot one? Thanks petulkacat@gmail.com
hi, I'm also currently facing the same challenge. Could you send me the conversion sheet of Sol gene IDs to UniProt IDs? my email: qiangli@uga.edu. Thanks
I got an excel sheet with EntrezGENEid and EnsembleGENEid (Solyc). If you are interested in this file, let me know.
Regards,
You can post the file up on pastebin.com or as a gist on github.com. Then post the link in your answer from yesterday.
Sorry, but the file is too big for pastebin.com
hey, could you please send me this excel sheet? you need my email or something? thanks
I am looking for this gene LOC101249187 ( NCBI Gene ID ) into the SolycXXgXXXXXX system
anyone can help ??